Pointers at Glance
- A new method, SPOTS, described in a paper in Nature Biotechnology, records gene activity patterns.
- SPOTS method can map the spatial organization of cells within tissues and could have significant implications for basic and clinical research.
A team of researchers led by Dr. Dan Landau of Weill Cornell Medicine and Dr. Marlon Stoeckius of 10x Genomics have developed a new method known as Spatial PrOtein and Transcriptome Sequencing (SPOTS).
This new method can map the spatial organization of cells, including cell types, activities, and interactions within tissue samples. SPOTS allows for creating high-resolution maps of organ tissues, including diseased organs and tumors, which could be useful in basic and clinical research.
Potential of SPOTS Method
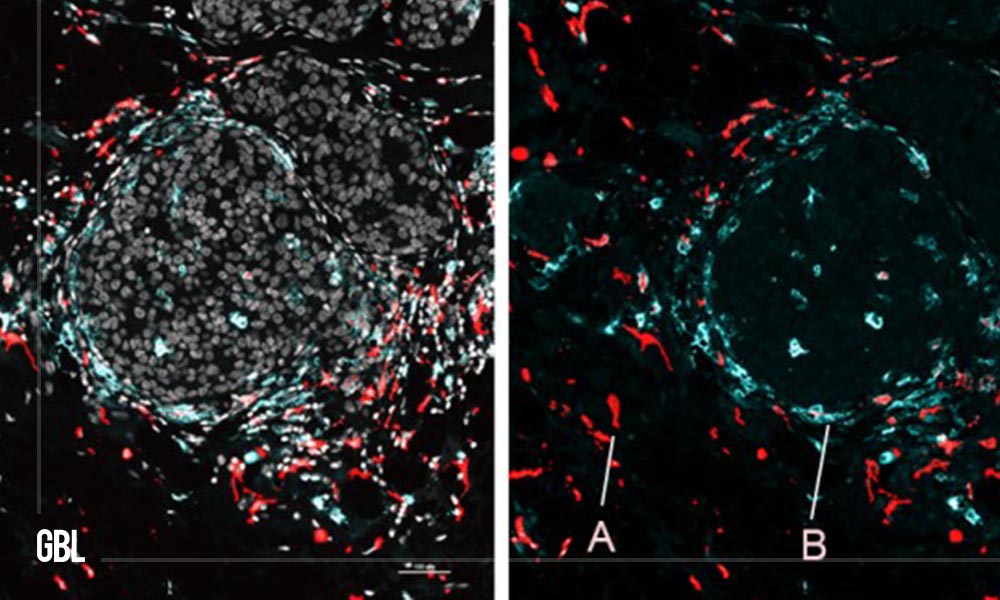
The team explained and highlighted the potential of SPOTS by using it to map the cellular organization of a mouse breast tumor and the functional architecture of a normal mouse spleen.
The SPOTS method is based on existing technology from 10x Genomics and uses glass slides that can be used for imaging tissue samples with conventional microscope-based pathology methods. These slides are coated with thousands of special probe molecules that contain a molecular “barcode” indicating their two-dimensional position on the slide.
When a thinly sliced tissue sample is placed on the slide, and the cells are made permeable, the probe molecules grab adjacent cells’ mRNAs, which are the transcripts of active genes.
Additionally, the method uses designer antibodies that bind to proteins of interest in the tissue and also bind to the special probe molecules. Using swift and automated techniques, researchers can identify the captured mRNAs and selected proteins and map them precisely to their original locations across the tissue sample.
Advantage of SPOTS Method
One of the key advantages of the SPOTS method is that it retains information about the cells’ precise locations within the tissue, which is often lost in other techniques for profiling gene activity and other layers of information in individual cells or small groups of cells. It allows for creating complex, data-rich “maps” of organs that can provide new insights into the spatial organization of cells within tissues.
Dr. Landau said that this initial version of SPOTS has a spatial resolution such that each “pixel” of the resulting dataset sums gene activity information for at least several cells.
The researchers hope to narrow this resolution to single cells soon while adding other layers of key cellular information.










hello!,I really like your writing very much! proportion we keep in touch more approximately your post on AOL? I require a specialist on this house to solve my problem. Maybe that’s you! Looking ahead to peer you.